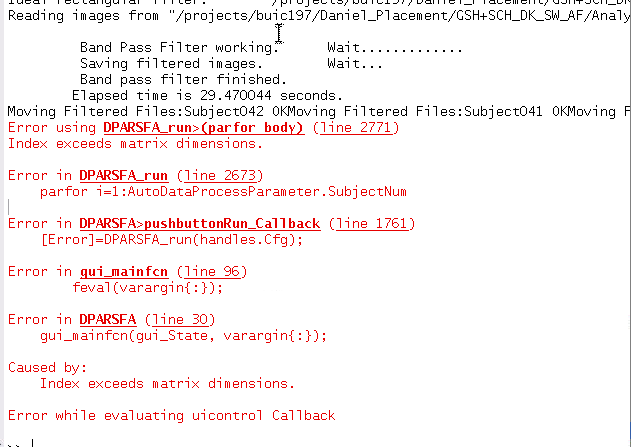
Run DPARSF from command line
- Read more about Run DPARSF from command line
- 2 comments
- Log in or register to post comments
After you select the options on the GUI and save the mat file is it possible to run DPARSF from the command line? The only machine I can run DPARSF that has the necessary Matlab toolboxes is on a linux cluster. I can easily submit jobs but am not permitted to run the GUI on the cluster. I am hoping that once the mat file is set kicking off the Analysis (i.e pushing the Run button) could be accomplished from the command line.
Thanks for your help.
Bob