The toolbox for Data Processing & Analysis for Brain Imaging on Surface (DPABISurf) (Yan et al., 2021) is based on fMRIPrep (Esteban et al., 2019), FreeSurfer (Dale et al., 1999), ANTs (Avants et al., 2008), FSL (Jenkinson et al., 2002), AFNI (Cox, 1996), SPM (Ashburner, 2012), dcm2niix (Li et al., 2016), PALM (Winkler et al., 2016), GNU Parallel (Tange, 2011), MATLAB (The MathWorks Inc., Natick, MA, US), Docker (https://docker.com) and DPABI (Yan et al., 2016). DPABISurf provides user-friendly graphical user interface (GUI) for pipeline surface-based preprocessing, statistical analyses and results viewing, while requires no programming/scripting skills from the users.
The DPABISurf pipeline first converts the user specified data into BIDS format (Gorgolewski et al., 2016), and then calls fMRIPprep docker to preprocess the structural and functional MRI data, which integrates FreeSurfer, ANTs, FSL and AFNI. With fMRIPprep, the data is processed into FreeSurfer fsaverage5 surface space and MNI volume space. DPABISurf further performs nuisance covariates regression (including ICA-AROMA) on the surface-based data (volume-based data is processed as well), and then calculate the commonly used R-fMRI metrics: amplitude of low frequency fluctuation (ALFF) (Zang et al., 2007), fractional ALFF (Zou et al., 2008), regional homogeneity (Zang et al., 2004), degree centrality (Zuo and Xing, 2014), and seed-based functional connectivity. DPABISurf also performs surface-based smoothing by calling FreeSurfer’s mri_surf2surf command. These processed metrics then enters surfaced-based statistical analyses within DPABISurf, which could perform surfaced-based permutation test with TFCE by integrating PALM. Finally, the corrected results could be viewed by the convenient surface viewer DPABISurf_VIEW, which is derived from spm_mesh_render.m.
DPABISurf is designed to make surface-based data analysis require minimum manual operations and almost no programming/scripting experience. We anticipate this open-source toolbox will assist novices and expert users alike and continue to support advancing R-fMRI methodology and its application to clinical translational studies.
DPABISurf is open-source and distributed under GNU/GPL, available with DPABI at http://www.rfmri.org/dpabi. It supports Windows 10 Pro, MacOS and Linux operating systems. You can run it with or without MATLAB.
1. With MATLAB.
1.1. Please go to http://www.rfmri.org/dpabi to download DPABI.
1.2. Add with subfolders for DPABI in MATLAB's path setting.
1.3. Input 'dpabi' and then follow the instructions of the "Install" Button on DPABISurf.
2. Without MATLAB.
2.1. Install Docker.
2.2. Terminal: docker pull cgyan/dpabi
2.3. Terminal: docker run -d --rm -v /My/FreeSurferLicense/Path/license.txt:/opt/freesurfer/license.txt -v /My/Data/Path:/data -p 5925:5925 cgyan/dpabi x11vnc -forever -shared -usepw -create -rfbport 5925
/My/FreeSurferLicense/Path/license.txt: Where you stored the FreeSurferLicense got from https://surfer.nmr.mgh.harvard.edu/registration.html.
/My/Data/Path: This is where you stored your data. In Docker, the path is /data.
2.4. Open VNC Viewer, connect to localhost:5925, the password is 'dpabi'.
2.5. In the terminal within the VNC Viewer, input "bash", and then input:
/opt/DPABI/DPABI_StandAlone/run_DPABI_StandAlone.sh ${MCRPath}
Now please enjoy the StandAlone version of DPABISurf with GUI!
References:
· Ashburner, J. (2012). SPM: a history. Neuroimage, 62(2), 791-800, doi:10.1016/j.neuroimage.2011.10.025.
· Avants, B.B., Epstein, C.L., Grossman, M., Gee, J.C. (2008). Symmetric diffeomorphic image registration with cross-correlation: evaluating automated labeling of elderly and neurodegenerative brain. Med Image Anal, 12(1), 26-41, doi:10.1016/j.media.2007.06.004.
· Cox, R.W. (1996). AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput Biomed Res, 29(3), 162-173.
· Dale, A.M., Fischl, B., Sereno, M.I. (1999). Cortical surface-based analysis. I. Segmentation and surface reconstruction. Neuroimage, 9(2), 179-194, doi:10.1006/nimg.1998.0395.
· Esteban, O., Markiewicz, C.J., Blair, R.W., Moodie, C.A., Isik, A.I., Erramuzpe, A., Kent, J.D., Goncalves, M., DuPre, E., Snyder, M., Oya, H., Ghosh, S.S., Wright, J., Durnez, J., Poldrack, R.A., Gorgolewski, K.J. (2019). fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat Methods, 16, 111-116, doi:10.1038/s41592-018-0235-4.
· Gorgolewski, K.J., Auer, T., Calhoun, V.D., Craddock, R.C., Das, S., Duff, E.P., Flandin, G., Ghosh, S.S., Glatard, T., Halchenko, Y.O., Handwerker, D.A., Hanke, M., Keator, D., Li, X., Michael, Z., Maumet, C., Nichols, B.N., Nichols, T.E., Pellman, J., Poline, J.B., Rokem, A., Schaefer, G., Sochat, V., Triplett, W., Turner, J.A., Varoquaux, G., Poldrack, R.A. (2016). The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci Data, 3, 160044, doi:10.1038/sdata.2016.44.
· Jenkinson, M., Bannister, P., Brady, M., Smith, S. (2002). Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage, 17(2), 825-841.
· Li, X., Morgan, P.S., Ashburner, J., Smith, J., Rorden, C. (2016). The first step for neuroimaging data analysis: DICOM to NIfTI conversion. J Neurosci Methods, 264, 47-56, doi:10.1016/j.jneumeth.2016.03.001.
· Tange, O. (2011). Gnu parallel-the command-line power tool. The USENIX Magazine, 36(1), 42-47.
· Winkler, A.M., Ridgway, G.R., Douaud, G., Nichols, T.E., Smith, S.M. (2016). Faster permutation inference in brain imaging. Neuroimage, 141, 502-516, doi:10.1016/j.neuroimage.2016.05.068.
· Yan, C.-G., Wang, X.-D., Lu, B. (2021). DPABISurf: data processing & analysis for brain imaging on surface. Sci Bull, 66(24), 2453-2455, doi: https://doi.org/10.1016/j.scib.2021.09.016.
· Yan, C.G., Wang, X.D., Zuo, X.N., Zang, Y.F. (2016). DPABI: Data Processing & Analysis for (Resting-State) Brain Imaging. Neuroinformatics, 14(3), 339-351, doi:10.1007/s12021-016-9299-4.
· Zang, Y., Jiang, T., Lu, Y., He, Y., Tian, L. (2004). Regional homogeneity approach to fMRI data analysis. Neuroimage, 22(1), 394-400, doi:http://dx.doi.org/10.1016/j.neuroimage.2003.12.030.
· Zang, Y.F., He, Y., Zhu, C.Z., Cao, Q.J., Sui, M.Q., Liang, M., Tian, L.X., Jiang, T.Z., Wang, Y.F. (2007). Altered baseline brain activity in children with ADHD revealed by resting-state functional MRI. Brain Dev, 29(2), 83-91, doi:10.1016/j.braindev.2006.07.002.
· Zou, Q.-H., Zhu, C.-Z., Yang, Y., Zuo, X.-N., Long, X.-Y., Cao, Q.-J., Wang, Y.-F., Zang, Y.-F. (2008). An improved approach to detection of amplitude of low-frequency fluctuation (ALFF) for resting-state fMRI: Fractional ALFF. Journal of Neuroscience Methods, 172(1), 137-141, doi:http://dx.doi.org/10.1016/j.jneumeth.2008.04.012.
· Zuo, X.-N., Xing, X.-X. (2014). Test-retest reliabilities of resting-state FMRI measurements in human brain functional connectomics: A systems neuroscience perspective. Neuroscience & Biobehavioral Reviews, 45, 100-118, doi:http://dx.doi.org/10.1016/j.neubiorev.2014.05.009.
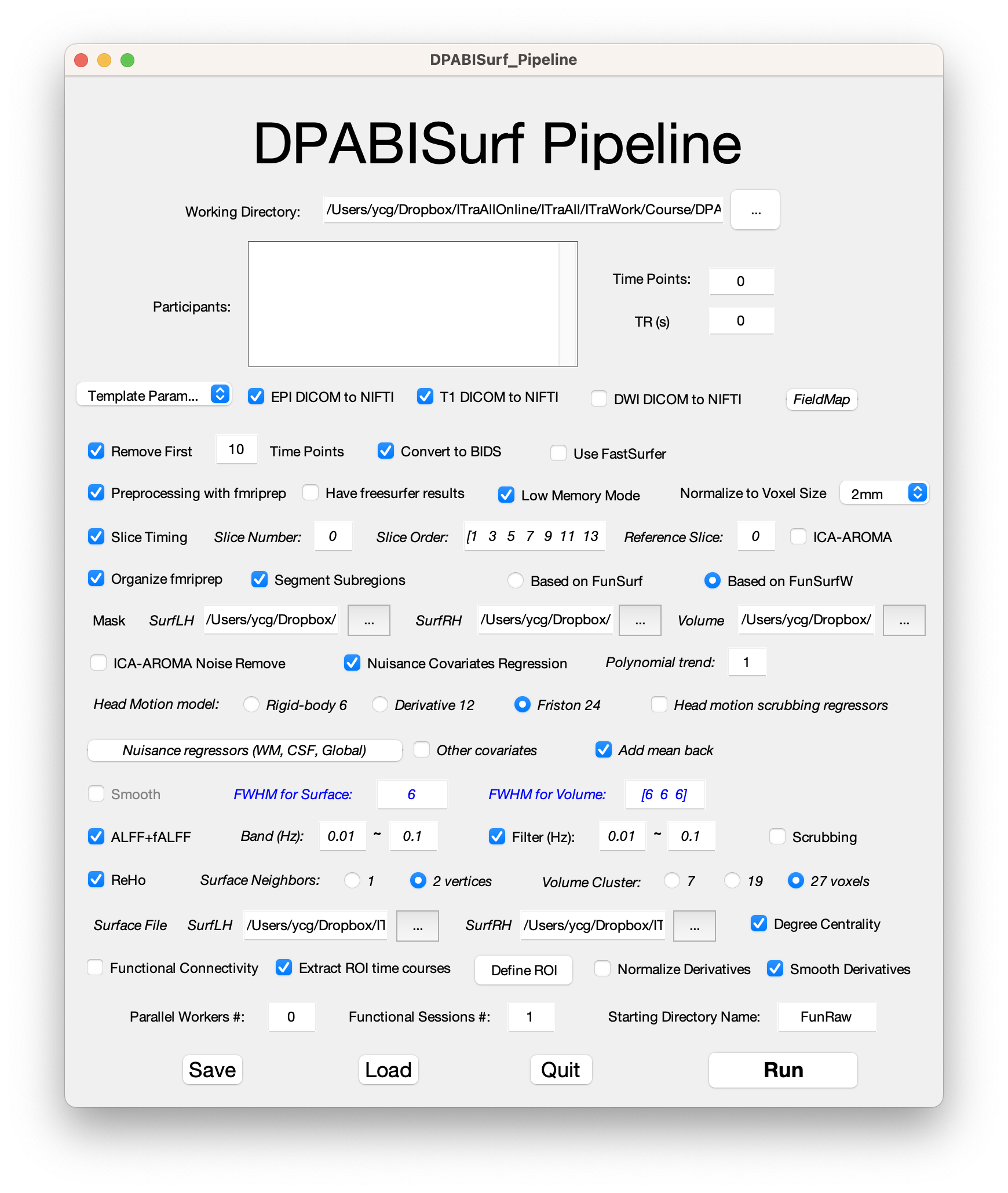


hippocampal tail's ROI
严老师您好,我想研究一下海马尾部和全脑的功能连接,但是不知道如何设置海马尾部的ROI,也找不到相关的mask,希望得到老师的帮助!
查查文献看freesurfer有这样的mask吗?
查查文献看freesurfer有这样的mask吗?
fmriprep: error Version 20.0.1 of fMRIPrep (current) FLAGGED
There is a minor bug when
There is a minor bug when using ICA-AROMA since fMRIPrep-20.0.1 changed the input format.
For "Version 20.0.1 of fMRIPrep (current) has been FLAGGED", you don't need to worry because DPABISurf doesn't use cifti-output.
As for now, please download our development version from https://github.com/Chaogan-Yan/DPABI
Please let me know if it works.
Thank you, with the GitHub
Thank you, with the GitHub version fMRIprep started running.
Matlab window has following error. Please let me know how to fix this. Thank you.
-------------------------------------------------------------
[Node] Running "lta2itk_fwd" ("niworkflows.interfaces.freesurfer.PatchedLTAConvert"), a CommandLine Interface with command:
lta_convert --inlta /data/fmriprepwork/sub-Sub002/fmriprep_wf/single_subject_Sub002_wf/anat_preproc_wf/surface_recon_wf/t1w2fsnative_xfm/out.lta --outitk /data/fmriprepwork/sub-Sub002/fmriprep_wf/single_subject_Sub002_wf/anat_preproc_wf/anat_derivatives_wf/lta2itk_fwd/out.txt
200327-06:36:15,538 nipype.workflow INFO:
[Node] Finished "fmriprep_wf.single_subject_Sub002_wf.anat_preproc_wf.anat_derivatives_wf.lta2itk_fwd".
200327-06:36:19,0 nipype.workflow ERROR:
could not run node: fmriprep_wf.single_subject_Sub002_wf.anat_preproc_wf.brain_extraction_wf.norm
You are using fMRIPrep-20.0.1, and a newer version of fMRIPrep is available: 20.0.5. Please check out our documentation about how and when to upgrade:
https://fmriprep.readthedocs.io/en/latest/faq.html#upgrading
WARNING: Version 20.0.1 of fMRIPrep (current) has been FLAGGED
(reason: fsLR resampling error (problematic only when using the --cifti-output flag)).
That means some severe flaw was found in it and we strongly
discourage its usage.
fMRIPrep failed: Workflow did not execute cleanly. Check log for details
Preprocessing did not finish successfully. Errors occurred while processing data from participants: Sub002 (1). Check the HTML reports for details.
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
Are you using demo data from:
Are you using demo data from: http://rfmri.org/demodata?
This is wierd, what's your OS?
Yes demo data from http:/
Yes demo data from http://rfmri.org/demodata
OS is MacOS Catalina 10.15.3
Thank you.
I don't have a Catalina to
I don't have a Catalina to test. But is should be OK.
Do you have a linux machine? Could you have a test on ubuntu or centos? Those platforms were tested.
dpabisur_run还是不能解决
老师好按照您说的,在github上下载了最新的工具包。仍然无法跑通这一步。用的是windows系统。
请问老师应该怎么办
resting-state 分析时,预处理报错
严老师,
您好!我在用dpabi做预处理的时候总是出现以下报错,想请问您是什么原因?应该怎么解决?
Seems you need to re-install
Seems you need to re-install SPM.
error in DPABISURF
严老师您好,
我在用DPABISurf进行到preprocessing with fmriprep过程中matlab突然被killed
我是在PALM的时候被killed,前面的都没问题
我是在PALM的时候被killed,前面的都没问题,就是到了PALM的时候,刚出了那个界面,就死了。
我是在PALM的时候被killed,前面的都没问题
我是在PALM的时候被killed,前面的都没问题,就是到了PALM的时候,刚出了那个界面,就死了。
DPABISurf 报错
严老师好!我用台式机windows10-pro系统(Intel i3-4160U 3.6GHz 双核4线程,16G内存,硬盘2T),Matlab2013b 运行DPABI_V4.3_200401中DPABISurf 1.3 【最新版docker2.3.0.3(45519),docker分了2个核,10G memory,load from local file <dpabi43_200401docker.tar.gz,licence.txt均顺利安装】,从DPABISurf_DemoData已做好fmriprep的步骤,照着视频设置往下跑(parallel:1#):
脑图维度转换
老师,您好,在处理静息态数据的时候,我想用自己的mask,但是我的mask是91x109x91,然后就没办法开展了,所以我想问一下如何将维度为91x109x91 的脑图转成61x73x61?
请使用dpabi>Utilities>Image
请使用dpabi>Utilities>Image Reslicer功能。
具体操作方法可以参考course中的演示。
好的,O(∩_∩)O谢谢
好的,O(∩_∩)O谢谢
使用dpabi分析HCP静息态数据分析
老师,您好,我从HCP数据库下载了他们预处理后的静息态数据,现在想用dpabi分析一下功能连接。问题是每个被试的静息态数据包括两个扫描方向的结果一个是从左向右扫描的静息态数据,如“rfMRI_REST1_LR”,一个是从右向左扫描的静息态数据“rfMRI_REST1_RL”。那如果使用dpabi分析的话,是先将这两种数据合并成一个数据吗?还是说先分开分析,最后将脑结果平均?麻烦了,期待回复!
分开计算然后平均可能更常用一些
分开计算然后平均可能更常用一些
HCP预处理后的公开数据跑功能连接报错
老师,您好,我使用HCP官网上预处理后的fMri数据(rfMRI_REST1_LR.nii)直接跑功能连接出现下面两张截图里的错误,这是什么原因导致的呢?

您好,我也使用HCP官网上预处理后的fMri数据
您好,我也使用HCP官网上预处理后的fMri数据,这种格式的(rfMRI_REST1_LR.nii)直接跑功能连接出现下面两张截图里的错误,这是为什么呢,您跑功能连接成功了么,怎么解决
HCP预处理后的公开数据跑功能连接报错
老师,您好,我使用HCP官网上预处理后的fMri数据(rfMRI_REST1_LR.nii)直接跑功能连接出现下面两张截图里的错误,这是什么原因导致的呢?

Do not add subfolders for
Do not add subfolders for REST and BrainNet Viewer.
Reorient这个过程完成之后会报错
严老师您好,我叫黄浦江
最近在使用dpabi 预处理T1的数据 Reorient这个过程完成之后会报错,想请问下您该如何解决,问题如下:
尝试在数据处理界面填入TR信息?
尝试在数据处理界面填入TR信息?
严老师您好:
严老师您好:
我在使用如下的afni指令进行图像的3D重构时,总是会遇到问题。
脚本指令:
uniq_images IMG*.dcm > uniq_image_list.txt
DPARSF预处理
老师您好:
我使用DPARSF进行任务态数据的预处理,发现一开始在FunRaw同样数量,同样大小的dcm文件,跑完预处理之后文件的大小在不同的run里不一样,大约差2mb,想请问一下发生这种情况的原因以及这样的数据会对后期的数据分析有影响吗?谢谢老师!
不影响。
不影响。
Dear, Prof. Yan
Dear, Prof. Yan
After re-run thefmriprep failed subjects, how should I choice the options?
'''
'''
best,
Hui Zheng
Seems these subjects failed
Seems these subjects failed fmriprep, please check.
严老师,dpabisurf分析时,在fmriprep的被试文件夹里面出现了log文件,没找到出现的问题,还请严老师帮我解决的一下
[environment]
cpu_count = 4
exec_env = "docker"
free_mem = 2.7
overcommit_policy = "heuristic"
overcommit_limit = "50%"
nipype_version = "1.5.1"
templateflow_version = "0.6.3"
version = "20.2.1"
[execution]
bids_dir = "/data/BIDS"
bids_database_dir = "/data/fmriprepwork/sub-sub0983/20210414-115444_7e921fc7-a77e-450c-8825-95139b8682d7/bids_db"
bids_description_hash = "ceb55f9ef8d5658e7418760e3be9884a4199d08a3fe8d1097cc0ad9faf5ae0a2"
boilerplate_only = false
sloppy = false
debug = []
fmriprep_dir = "/data/fmriprep"
fs_license_file = "/opt/freesurfer/license.txt"
fs_subjects_dir = "/data/freesurfer"
layout = "BIDS Layout: .../data/BIDS | Subjects: 1 | Sessions: 0 | Runs: 0"
log_dir = "/data/fmriprep/logs"
log_level = 25
low_mem = true
md_only_boilerplate = false
notrack = false
output_dir = "/data"
output_layout = "legacy"
output_spaces = "fsaverage:den-10k MNI152NLin2009cAsym:res-native"
reports_only = false
run_uuid = "20210414-115444_7e921fc7-a77e-450c-8825-95139b8682d7"
participant_label = [ "sub0983",]
templateflow_home = "/home/fmriprep/.cache/templateflow"
work_dir = "/data/fmriprepwork/sub-sub0983"
write_graph = false
[workflow]
anat_only = false
aroma_err_on_warn = false
aroma_melodic_dim = -200
bold2t1w_dof = 6
bold2t1w_init = "register"
cifti_output = false
fmap_bspline = false
force_syn = false
hires = true
ignore = []
longitudinal = false
medial_surface_nan = false
regressors_all_comps = false
regressors_dvars_th = 1.5
regressors_fd_th = 0.5
run_reconall = true
skull_strip_fixed_seed = false
skull_strip_template = "OASIS30ANTs"
skull_strip_t1w = "force"
spaces = "fsaverage:den-10k MNI152NLin2009cAsym:res-native"
use_aroma = false
use_syn_sdc = false
[nipype]
crashfile_format = "txt"
get_linked_libs = false
nprocs = 4
omp_nthreads = 3
plugin = "MultiProc"
resource_monitor = true
stop_on_first_crash = false
[seeds]
master = 536
ants = 25065
[nipype.plugin_args]
maxtasksperchild = 1
raise_insufficient = false
这个log文件不是报错文件
这个log文件不是报错文件。DPABISurf应该能继续跑下去。
Troubleshooting
Hello,
I am new to DPABI Surf and wanted to use its ICA-AROMA noise remove method for my own analysis. I have installed the Docker as it described in the website and got the varification for it as well. However I still get error which I hope someone here can help me fixing it. I got the following lines of errors:
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/rh.area
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ lh.curv
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ rh.curv
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ lh.sulc
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ rh.sulc
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ lh.volume
ERROR: could not determine type of /data/freesurfer/sub-NV163V1/surf/ rh.volume
Error using y_Organize_fmriprep>(parfor body) (line 118)
cp: cannot stat
'/mnt/MXMVPGroupShare/JXH108/NV_Sanders/Original_Data/NoDelyn/NoDelyn_12_16_2020/fmriprep/sub-NV004V1/anat/*_space-MNI152NLin2009cAsym_*':
No such file or directory
Error in y_Organize_fmriprep (line 116)
parfor i=1:Cfg.SubjectNum
Error in DPABISurf_run (line 479)
y_Organize_fmriprep(Cfg);
Error in DPABISurf_Pipeline>pushbuttonDPABISurfRun_Callback (line 1693)
[Error, Cfg]=DPABISurf_run(handles.Cfg);
Error in gui_mainfcn (line 95)
feval(varargin{:});
Error in DPABISurf_Pipeline (line 42)
gui_mainfcn(gui_State, varargin{:});
Error in
@(hObject,eventdata)DPABISurf_Pipeline('pushbuttonDPABISurfRun_Callback',hObject,eventdata,guidata(hObject))
Error while evaluating UIControl Callback
Thanks in advance for your helps!
The running of fmriprep
The running of fmriprep failed. Please check.
预处理时候遇到的错误
老师好!
我在预处理的时候点完reorient的图像后,便会出现以下报错,请问该怎么解决呢?谢谢老师!
通常是数据整理的问题。具体原因从目前信息无法判断。
通常是数据整理的问题。具体原因从目前信息无法判断。
严老师您好,我在网上下载了其他的数据想进行数据处理
严老师您好,我在网上下载了其他的数据想进行数据处理,但是跑了很多次都在同一个地方报错,我自己不太可能解决这个问题,希望得到老师的帮助!以下是报错的内容
看起来是这个数据有问题。你可以先试试别的数据。
看起来是这个数据有问题。你可以先试试别的数据。
DPABISurf报错
严老师您好,我最近跑DPABISurf会出现报错,有几个地方我觉得很疑惑
(1)fmriprep这一步被报告没跑成功的被试在其对应的网页上却显示:
你试试demo数据也这样吗?
你试试demo数据也这样吗?
老师您好!
老师您好!
请参阅:http://www.rfmri.org
请参阅:http://www.rfmri.org/SliceTiming
Siemens 3.0T:Field map correction
严老师您好:
用西门子3.0T核磁采集的fmri序列,同时采集了bold_filed_maping序列,如果想做Field map correction,原始数据该怎么设置?
您的示范数据—FieldMap中放了3个文件夹,谢谢!
请参阅:
请参阅:
https://github.com/Puyunfashi/Suzhou_rumination/blob/master/sort_data/Organize_FieldMap.m
和
https://github.com/Puyunfashi/Suzhou_rumination/blob/master/sort_data/c_DivideFieldMapMagnitudeFiles.m
修改了标准的ICBM/MNI脑模板后再做new segment+dartel得不到正确的分割和标准化结果
严老师:
您好!
我目前在做一组儿童的静息态fMRI预处理,因此在https://www.nitrc.org/projects/chn-pd这里下载了贺永老师团队创建的一个中国儿童标准脑模板,利用其TPM先验模板来做segment和normalize,但得到的灰质、白质分割结果是一个这样的图,因此我想请问老师为什么会出现这样的结果,有什么方法可以解决吗?如果我做更换模板这样的操作,需要注意哪些问题呢?
很明显分割有问题。
很明显分割有问题。
要替换DPABI/Templates/SPMTemplates/tpm/TPM.nii
严老师,您好!
严老师,您好!
以上这个结果是我做了替换后得到的分割结果,而且同时也用spm尝试了一下,发现也是这样的,目前没有找到问题出在哪,麻烦老师帮忙解答一下,谢谢!
DPABIsurf
严老师您好,我有个问题想请教您,您在DPABISurf输出文件夹与计算结果概述 的课程里提到-----存在Data>fmriprep>sub-Sub001>log文件夹的话,一般是跑fmriprep可能出问题了。老师我想问您,我还要继续运行DPABIsurf吗?
最新的fmriprep更改了逻辑
最新的fmriprep更改了逻辑,有log文件夹不一定代表出错。DPABISurf会自己判断,如果不报错就是正常。