 |
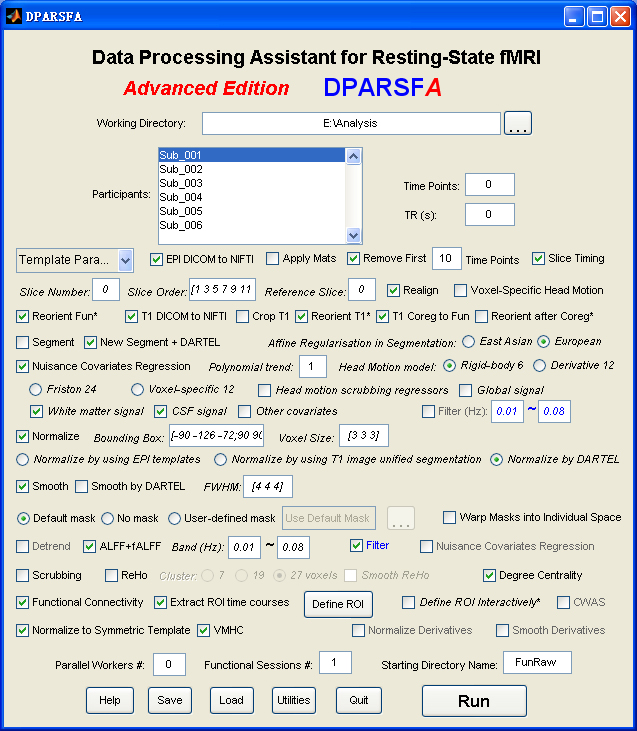
Data Processing Assistant for Resting-State fMRI (DPARSF) is a convenient plug-in software based on SPM and REST. You just need to arrange your DICOM files, and click a few buttons to set parameters, DPARSF will then give all the preprocessed (slice timing, realign, normalize, smooth) data, functional connectivity, ReHo, ALFF/fALFF, degree centrality, voxel-mirrored homotopic connectivity (VMHC) results. DPARSF can also create a report for excluding subjects with excessive head motion and generate a set of pictures for easily checking the effect of normalization. You can use DPARSF to extract ROI time courses efficiently if you want to perform small-world analysis. DPARSF basic edition is very easy to use while DPARSF advanced edition (alias: DPARSFA) is much more flexible and powerful. DPARSFA can parallel the computation for each subject, and can be used to reorient your images interactively or define regions of interest interactively. You can skip or combine the processing steps in DPARSF advanced edition freely. Please download a MULTIMEDIA COURSEto know more about how to use this software. Add DPARSF's directory to MATLAB's path and enter "DPARSF" or "DPARSFA" in the command window to enjoy DPARSF basic edition or advanced edition. The latest release is DPARSF_V2.3_130615. |
New features of DPARSF_V2.3_130615:
1. Apply downloaded reorient matrices. Given the amount of time and effort for interactive reorienting, the reorient matrices for online data such as 1000 Functional Connectomes Project (FCP) (Biswal et al., 2010) and Autism Brain Imaging Data Exchange (ABIDE) (Di Martino et al., 2013) could be downloaded and applied automatically. Please find here for “DownloadedReorientMats”, and then tick the checkbox "Apply Mats".
2. Save information of TR, Slice Number, Time Points and Voxel Size into TRInfo.tsv (under working directory) file for checking data correctness.
3. The slice order type could be specified for each participant into SliceOrderInfo.tsv (under working directory) file, thus allow different slice timing correction for different participants in a batch mode. Please find instructions for setting SliceOrderInfo.tsv from {DPARSF}/Docs/SliceOrderInfo.tsv_Instruction.txt.
4. The output format of DICOM to NIfTI were changed to 4D .nii images.
5. The midline of VMHC results were set to zero.
Biswal, B.B., Mennes, M., Zuo, X.N., Gohel, S., Kelly, C., Smith, S.M., Beckmann, C.F., Adelstein, J.S., Buckner, R.L., Colcombe, S., Dogonowski, A.M., Ernst, M., Fair, D., Hampson, M., Hoptman, M.J., Hyde, J.S., Kiviniemi, V.J., Kotter, R., Li, S.J., Lin, C.P., Lowe, M.J., Mackay, C., Madden, D.J., Madsen, K.H., Margulies, D.S., Mayberg, H.S., McMahon, K., Monk, C.S., Mostofsky, S.H., Nagel, B.J., Pekar, J.J., Peltier, S.J., Petersen, S.E., Riedl, V., Rombouts, S.A., Rypma, B., Schlaggar, B.L., Schmidt, S., Seidler, R.D., G, J.S., Sorg, C., Teng, G.J., Veijola, J., Villringer, A., Walter, M., Wang, L., Weng, X.C., Whitfield-Gabrieli, S., Williamson, P., Windischberger, C., Zang, Y.F., Zhang, H.Y., Castellanos, F.X., Milham, M.P., 2010. Toward discovery science of human brain function. Proc Natl Acad Sci U S A 107, 4734-4739.
Di Martino, A., Yan, C.G., Li, Q., Denio, E., Castellanos, F.X., Alaerts, K., Anderson, J.S., Assaf, M., Bookheimer, S.Y., Dapretto, M., Deen, B., Delmonte, S., Dinstein, I., Ertl-Wagner, B., Fair, D.A., Gallagher, L., Kennedy, D.P., Keown, C.L., Keysers, C., Lainhart, J.E., Lord, C., Luna, B., Menon, V., Minshew, N., Monk, C.S., Mueller, S., Muller, R.A., Nebel, M.B., Nigg, J.T., O’Hearn, K., Pelphrey, K.A., Peltier, S.J., Rudie, J.D., Sunaert, S., Thioux, M., Tyszka, J.M., Uddin, L.Q., Verhoeven, J.S., Wenderoth, N., Wiggins, J.L., Mostofsky, S.H., Milham, M.P., 2013. The Autism Brain Imaging Data Exchange: towards large-scale evaluation of the intrinsic brain architecture in Autism. Mol Psychiatry in press.
New features of DPARSF_V2.2_130309:
1. Advanced Edition: Fixed a bug in "Smooth by DARTEL" caused in previous revision (DPARSF_V2.2_130214). (Thanks for the report of Maki Koyama).
New features of DPARSF_V2.2_130303:
1. Advanced Edition: Fixed a bug in processing of multiple sessions - only process the last session in normalization and smooth.
2. Basic Edition: Fixed a bug caused in the previous release (V2.2_130214) - cannot load the correct masks (Thanks for the report of Tao Yang).
New features of DPARSF_V2.2_130224:
1. This release fixed some minor bugs, but will not affect the results of any data analysis. The bugs appear in uncommon parameter settings and stop the processing in the worst cases.
2. Updated the DPARSFVersion information in the template parameter files.
3. DPARSF basic edition will also output the results of ReHo/ALFF/fALFF after Z-standardization (subtract the whole brain mean and divide by the whole brain standard deviation).
4. Fixed a bug in nuisance covariates regression when CovMat is not defined.
5. Fixed a bug that can not save the ROI signals correctly in text file if the data is not in double format. (Thanks for the report by H. Baetschmann)
6. Fixed a bug in creating mean functional image for 4D files. (Thanks for the report and revision by S. Orsolini)
7. Fixed a bug in displaying ROI Templates in linux system. (Thanks for the report by Han Zhang)
New features of DPARSF_V2.2_121225:
1. Support parallel computing! If you installed the MATLAB parallel computing toolbox, you can set the number of "Parallel Workers", DPARSFA will distribute the subjects into different CPU cores.
2. In addressing head motion concerns in resting-state fMRI analyses (Power et al., 2012; Satterthwaite et al., 2012; Van Dijk et al., 2012), we provide Friston 24-parameter correction (See Yan et al., 2013 for a comprehensive assessment of head motion on functional connectomics) as well as voxel-specific head motion calculation and correction (Satterthwaite et al., 2013; Yan et al., 2013). DPARSF also calculate the voxel-specific mean framewise displacement (FD) and volume-level mean FD (Power) (Power et al., 2012) or FD (Jenkinson) (i.e., relative RMS; Jenkinson et al., 2002) for accounting head motion at group-level analysis. The data scrubbing approach is also supported with different methods: 1) model each bad time point as a separate regressor in nuisance covariates regression, 2) delete bad time points, 3) interpolate bad time points with nearest neighbor, linear or cubic spline interpolation.
3. According to (Weissenbacher et al., 2009), the nuisance covariate regression could be performed before filtering and at very early stage. Users can also choose Template Parameters: TRADITIONAL order to have the same order as the previous version.
4. Support .nii.gz in all the steps. No longer need to convert 4D .nii.gz into 3D .img/.hdr. Simply put .nii.gz under FunImg or any starting directory name, DPARSFA will handle the .nii.gz by itself.
5. If the Number of Time Points is set to 0, then DPARSFA will not check the number of time points.
6. If TR is set to 0, then DPARSFA will retrieve the TR information from the NIfTI images. Please ensure the TR information in NIfTI images are correct!
7. If Slice Number is set to 0, then retrieve the slice number from the NIfTI images. The slice order is then assumed as interleaved scanning: [1:2:SliceNumber, 2:2:SliceNumber]. The reference slice is set to the slice acquired at the middle time point, i.e., ceil(SliceNumber/2). SHOULD BE EXTREMELY CAUTIOUS!!!
8. Support calculating resting-state fMRI metrics in native space and warping by DARTEL. The mask files and/or region of interest (ROI) files in standard space are warped into native space by using the parameters estimated in segmentation or DARTEL.
9. Spatial normalization and smooth can be performed on the calculated resting-state fMRI derivatives.
10. Supporting extracting ROI time course using masks with multiple labels. More template ROI definitions are supported: Dosenbach’s 160 functional ROIs (Dosenbach et al., 2010), Andrews-Hanna’s default mode network ROIs (Andrews-Hanna et al., 2010), Craddock’s clustering ROIs (Craddock et al., 2011), AAL atlas and Harvard-Oxford atlas.
11. More resting-state fMRI metrics are included, e.g., voxel-mirrored homotopic connectivity (VMHC) (Zuo et al., 2010), Degree Centrality (Buckner et al., 2009) and connectome-wide association studies based on multivariate distance matrix regression (Shehzad et al., 2011).
12. For VMHC analyses, support normalizing the data further to a symmetric template. 1) Get the T1 images in MNI space (e.g., wco*.img or wco*.nii under T1ImgNewSegment or T1ImgSegment) for each subject, and then create a mean T1 image template (averaged across all the subjects). 2) Create a symmetric T1 template by averaging the mean T1 template (created in Step 1) with it's flipped version (flipped over x axis). 3) Normalize the T1 image in MNI space (e.g., wco*.img or wco*.nii under T1ImgNewSegment or T1ImgSegment) for each subject to the symmetric T1 template (created in Step 2), and apply the transformations to the functional data (which have been normalized to MNI space beforehand). Please see a reference from Zuo et al., 2010.
13. A group analyses function was added as y_GroupAnalysis_Image by scripting call. T tests or F tests could be performed for a given set of regressors.
14. Template Parameters:
Calculate in Original Space (warp by DARTEL)
Calculate in Original Space (warp by information from unified segmentation)
Calculate in MNI Space (warp by DARTEL)
Calculate in MNI Space (warp by information from unified segmentation)
Calculate in MNI Space: TRADITIONAL order [This is the order used in DPARSF basie edition as well as DPARSFA V2.1]
Intraoperative Processing
Task fMRI data preprocessing
VBM (New Segment and DARTEL)
VBM (unified segmentation)
Blank
Many thanks to Dr. Chris Rorden for suggesting features 4-7. Many thanks to Dr. Susan Whitfield-Gabrieli for discussing the head motion scrubbing regressors.
References:
Andrews-Hanna, J.R., Reidler, J.S., Sepulcre, J., Poulin, R., Buckner, R.L., 2010. Functional-anatomic fractionation of the brain's default network. Neuron 65, 550-562.
Buckner, R.L., Sepulcre, J., Talukdar, T., Krienen, F.M., Liu, H., Hedden, T., Andrews-Hanna, J.R., Sperling, R.A., Johnson, K.A., 2009. Cortical hubs revealed by intrinsic functional connectivity: mapping, assessment of stability, and relation to Alzheimer's disease. J Neurosci 29, 1860-1873.
Craddock, R.C., James, G.A., Holtzheimer, P.E., 3rd, Hu, X.P., Mayberg, H.S., 2011. A whole brain fMRI atlas generated via spatially constrained spectral clustering. Hum Brain Mapp.
Dosenbach, N.U., Nardos, B., Cohen, A.L., Fair, D.A., Power, J.D., Church, J.A., Nelson, S.M., Wig, G.S., Vogel, A.C., Lessov-Schlaggar, C.N., Barnes, K.A., Dubis, J.W., Feczko, E., Coalson, R.S., Pruett, J.R., Jr., Barch, D.M., Petersen, S.E., Schlaggar, B.L., 2010. Prediction of individual brain maturity using fMRI. Science 329, 1358-1361.
Jenkinson, M., Bannister, P., Brady, M., Smith, S., 2002. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17, 825-841.
Power, J.D., Barnes, K.A., Snyder, A.Z., Schlaggar, B.L., Petersen, S.E., 2012. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142-2154.
Satterthwaite, T.D., Wolf, D.H., Loughead, J., Ruparel, K., Elliott, M.A., Hakonarson, H., Gur, R.C., Gur, R.E., 2012. Impact of in-scanner head motion on multiple measures of functional connectivity: Relevance for studies of neurodevelopment in youth. Neuroimage 60, 623-632.
Satterthwaite, T.D., Elliott, M.A., Gerraty, R.T., Ruparel, K., Loughead, J., Calkins, M.E., Eickhoff, S.B., Hakonarson, H., Gur, R.C., Gur, R.E., Wolf, D.H., 2013. An Improved Framework for Confound Regression and Filtering for Control of Motion Artifact in the Preprocessing of Resting-State Functional Connectivity Data. Neuroimage 64, 240-256.
Shehzad, Z., Reiss, P.T., Adelstein, J., Emerson, J.W., Chabernaud, C., Mennes, M., DiMartino, A., McMahon, K., Copland, D., Castellanos, F.X., Kelly, C., Milham, M.P., 2011. Connectome-Wide Association Studies (CWAS). 17th Annual Meeting of the Organization for Human Brain Mapping, Quebec City.
Van Dijk, K.R., Sabuncu, M.R., Buckner, R.L., 2012. The influence of head motion on intrinsic functional connectivity MRI. Neuroimage 59, 431-438.
Weissenbacher, A., Kasess, C., Gerstl, F., Lanzenberger, R., Moser, E., Windischberger, C., 2009. Correlations and anticorrelations in resting-state functional connectivity MRI: a quantitative comparison of preprocessing strategies. Neuroimage 47, 1408-1416.
Yan, C.G., Cheung, B., Kelly, C., Colcombe, S., Craddock, R.C., Di Martino, A., Li, Q., Zuo, X.N., Castellanos, F.X., Milham, M.P., 2013. A comprehensive assessment of regional variation in the impact of head micromovements on functional connectomics. Neuroimage 76, 183-201.
Zuo, X.N., Kelly, C., Di Martino, A., Mennes, M., Margulies, D.S., Bangaru, S., Grzadzinski, R., Evans, A.C., Zang, Y.F., Castellanos, F.X., Milham, M.P., 2010. Growing together and growing apart: regional and sex differences in the lifespan developmental trajectories of functional homotopy. J Neurosci 30, 15034-15043.
New features of DPARSF_V2.1_120101:
For DPARSFA (Advanced Edition):
1. Support .nii and .nii.gz 3D or 4D files. For 4D .nii(.gz) functional files, use Checkbox "4D Fun .nii(.gz) to 3D" to convert into 3D files. For T1 3D .nii.gz files, use Checkbox "Unzip T1 .gz" to unzip. Use Checkbox "Crop T1" to Reorient to the nearest orthogonal direction to "canonical space" and remove excess air surrounding the individual as well as parts of the neck below the cerebellum (MRIcroN's dcm2nii).
2. Normalize by DARTEL has been added. Details: (1) "T1 Coreg to Fun": the individual structural T1 image is coregistered to the mean functional image after motion correction. (2) "New Segment + DARTEL": New Segment -- The transformed structural image is then segmented into gray matter, white matter and cerebrospinal fluid by using "New Segment" in SPM8. (3) "New Segment + DARTEL": DARTEL -- Create Template, and DARTEL -- Normalize to MNI space (Many Subjects) for GM, WM, CSF and T1 Images (unmodulated, modulated and smoothed [8 8 8] kernel versions). (4) "Normalize by DARTEL": DARTEL Normalize to MNI space (Few Subjects) for functional images. (5) "Smooth by DARTEL": DARTEL Normalize to MNI space (Few Subjects) for functinal images but with smooth kernel as specified, the smoothing is part of the normalisation to MNI space computes these average intensities from the original data, rather than the warped versions.
3. Reorient functional images and reorient T1 images interactively before coregistration: Checkbox "Reorient Fun*" and Checkbox "Reorient T1*". Interactively reorienting the anatomic images and functional images so that the origin approximated the anterior commissure and the orientation approximated MNI space, this will improve the accuracy in coregistration and segmentation. This step could probably solve the bad normalization problem for some subjects in "normalized by unified segmentation" or "normalized by DARTEL".
4. Multiple functional sessions supported. The directory should be named as FunRaw (or FunImg) for the first session; S2_FunRaw (or S2_FunImg) for the second session; and S3_FunRaw (or S3_FunImg) for the third session... In "Realign", "the sessions are first realigned to each other, by aligning the first scan from each session to the first scan of the first session. Then the images within each session are aligned to the first image of the session." (from SPM Manual).
5. Fixed a bug for calculation error in the second (and 3rd, 4th, ...) subjects in "Calculate in Original Space (Warp by information in unified segmentation)".
6. The calculations of ALFF and fALFF are promoted before filtering. Fixed a previous bug of calculating fALFF after filtering in the previous version of DPARSFA.
7. Mac OS compatible.
8. Template Parameters in DPARSFA:
8.1. Standard Steps: Normalized by DARTEL
8.2. Standard Steps: Normalized by DARTEL (Start from .nii.gz files)
8.3. Standard Steps: Normalized by T1 image unified segmentation
8.4. Calculate in Original Space (Warp by information in unified segmentation)
8.5. Intraoperative Processing
8.6. VBM (New Segment and DARTEL)
8.7. VBM (unified segmentaition)
8.8. Blank
For DPARSF (Basic Edition)
1. Normalize by DARTEL has been added. By checking "Normalized by using.. DARTEL", the processing details are the same as in DPARSFA: (1) "T1 Coreg to Fun": the individual structural T1 image is coregistered to the mean functional image after motion correction. (2) "New Segment + DARTEL": New Segment -- The transformed structural image is then segmented into gray matter, white matter and cerebrospinal fluid by using "New Segment" in SPM8. (3) "New Segment + DARTEL": DARTEL -- Create Template, and DARTEL -- Normalize to MNI space (Many Subjects) for GM, WM, CSF and T1 Images (unmodulated, modulated and smoothed [8 8 8] kernel versions). (4) "Normalize by DARTEL": DARTEL Normalize to MNI space (Few Subjects) for functional images. (5) "Smooth by DARTEL": DARTEL Normalize to MNI space (Few Subjects) for functinal images but with smooth kernel as specified, the smoothing is part of the normalisation to MNI space computes these average intensities from the original data, rather than the warped versions.
Hope to finish a video course for the new features in soon.
New features of DPARSF_V2.0_110505:
1. Fixed an error in the future MATLAB version in "[pathstr, name, ext, versn] = fileparts...".
New features of DPARSF_V2.0_101025:
1. DPARSF advanced edition (alias: DPARSFA) is added with the following new features:
1.1. The processing steps can be freely skipped or combined.
1.2. The processing can be start with any Starting Directory Name.
1.3. Support ReHo, ALFF/fALFF and Functional Connectivity calculation in individual space.
1.4. The masks or ROI files would be resampled automatically if the dimension mismatched the functional images.
1.5. The masks or ROI files in standard space can be warped into individual space by using the parameters estimated in unified segmentaion.
1.6. Support VBM analysis by checking "Segment" only.
1.7. Support reorientation interactively if the images in a bad orientation.
1.8. Support define regions of interest interactively based on the participant's T1 image in individual space.
2. DPARSF basic edition is preserved with the same operation style with DPARSF V1.0. DPARSF basic edition has the following new features:
2.1. Fixed a bug in copying "*.ps" files.
2.2. Will not check "wra*" prefix in "FunImgNormalized" directory.
2.3. Fixed a bug while regress out head motion parameters only.
The multimedia course for DPARSF advanced edition is estimated to be released in this November, thanks for your patience.
New features of DPARSF_V1.0_100510:
1. Added a right-click menu to delete all the participants' ID.
2. Fixed a bug in converting DICOM files to NIfTI in Windows 7, thanks to Prof. Chris Rorden's new dcm2nii.
3. Now will detect if co* T1 image (T1 image which is reoriented to the nearest orthogonal direction to 'canonical space' and removed excess air surrounding the individual as well as parts of the neck below the cerebellum) exists before normalization by using T1 image unified segmentation. T1 image without 'co' is also allowed in the analysis now.
New features of DPARSF_V1.0_100420:
1. After extracting ROI time courses, not just functional connectivity will be calculated, but also transform the r values to z values by Fisher's z transformation.
2. Fixed a bug in generating pictures for checking normalization when the bounding box is not [-90 -126 -72;90 90 108].
New features of DPARSF_V1.0_100201:
1. Save the configuration parameters automatically.
2. Fixed the bug in converting DICOM files to NIfTI files when DPARSF stored under C:\Program Files\Matlab\Toolbox.
3. Fixed the bug in converting DICOM files to NIfTI files when the filename without extension.
New features of DPARSF_V1.0_091215:
1. Also can regress out other kind of covariates other than head motion parameters, Global mean signal, White matter signal and Cerebrospinal fluid signal.
New features of DPARSF_V1.0_091201:
1. Added an option to choose different Affine Regularisation in Segmentation: East Asian brains (eastern) or European brains (mni). The interpretation of this option from SPM is: “If you can approximately align your images prior to running Segment, then this will increase the robustness of segmentation. Another thing that may help would be to change the regularisation of the initial affine registration, via Segment->Custom->Affine Regularisation. If you set this to "ICBM space template - East Asian brains (or European brains)", then the algorithm will make use of knowledge about the approximate variability to expect among the width/length etc of the brains of the population.” “The prior probability distribution for affine registration of East-Asian brains to MNI space was derived from 65 seg_inv_sn.mat files from Singapore. The distribution of affine transforms of European brains was estimated from: Incorporating Prior Knowledge into Image Registration NeuroImage, Volume 6, Issue 4, November 1997, Pages 344-352 J. Ashburner, P. Neelin, D. L. Collins, A. Evans, K. Friston.”
2. Added a Utility: change the Prefix of Images since DPARSF need some special prefixes in some cases. For example, if you do not have T1 DICOM files and your T1 NIFTI files are not initiated with “co”, then you can use this utility to add the “co” prefix to let DPARSF perform normalization based on segmentation of T1 images.
3. Added a popup menu to delete selected subject by right click.
4. Added a checkbox for removing first time points.
5. Added a function to close wait bar when program finished.
New features of DPARSF_V1.0Beta_091001:
1. SPM8 compatible.
2. Generate the pictures (output in {Working Directory}\PicturesForChkNormalization\) for checking normalization.
New features of DPARSF_V1.0Beta_090911:
1. Fixed the bug of setting user's defined mask.
New features of DPARSF_V1.0Beta_090901:
1. Fixed the bug of setting FWHM kernel of smooth.
2. Smooth the mReHo results.
3. Remove any number of the first time points.
New features of DPARSF_V1.0Beta_090713:
1. mReHo - 1, mALFF - 1, mfALFF - 1 function.
2. Creating report for excessive head motion subjects excluding.
New features of DPARSF_V1.0Beta_090701:
1. Linux compatible.
DPARSF's standard processing steps:
1. Convert DICOM files to NIFTI images.
2. Remove First 10 (more or less) Time Points.
3. Slice Timing.
4. Realign.
5. Normalize.
6. Smooth (optional).
7. Detrend.
8. Filter.
9. Calculate ReHo, ALFF, fALFF (optional).
10. Regress out the Covariables (optional).
11. Calculate Functional Connectivity (optional).
12. Extract AAL or ROI time courses for further analysis (optional).
-----------------------------------------------------------
Citing Information:
If you think DPARSFA is useful for your work, citing it in your paper would be greatly appreciated.
Something like "... The preprocessing was carried out by using Data Processing Assistant for Resting-State fMRI (DPARSF) (Yan & Zang, 2010, http://www.restfmri.net) which is based on Statistical Parametric Mapping (SPM8) (http://www.fil.ion.ucl.ac.uk/spm) and Resting-State fMRI Data Analysis Toolkit (REST, Song et al., 2011. http://www.restfmri.net)..."
Reference: Yan C and Zang Y (2010) DPARSF: a MATLAB toolbox for "pipeline" data analysis of resting-state fMRI. Front. Syst. Neurosci. 4:13. doi:10.3389/fnsys.2010.00013; Song, X.W., Dong, Z.Y., Long, X.Y., Li, S.F., Zuo, X.N., Zhu, C.Z., He, Y., Yan, C.G., Zang, Y.F., 2011. REST: A Toolkit for Resting-State Functional Magnetic Resonance Imaging Data Processing. PLoS ONE 6, e25031.
DPARSF is based on MRIcroN' dcm2nii, SPM and REST, if you used the related modules, the following software may need to be cited:
Step 1: MRIcroN software (by Chris Rorden, http://www.mricro.com).
Step 3 - Step 6: Statistical Parametric Mapping (SPM8, http://www.fil.ion.ucl.ac.uk/spm).
Step 7 - Step 11: Resting-State fMRI Data Analysis Toolkit (REST, Song et al., 2011. http://www.restfmri.net)
Licence: GNU General Public Licence (GPL)

DPARSFA怎样导入数据
严老师:
您好!我最近在学习DPARSFA,但一开始就遇到了问题,我将数据文件夹FunImg的上一级文件夹作为数据路径放入MATLAB,但是在DPARSFA中无法导入数据,DPARSFA和DPARSF放数据的路径Working Directory设置不一样吗?DPARSFA应该怎么设置Working Directory呢?
望老师能给与指导,谢谢老师!
Hi,
Hi,
You'll see the subjects after you setting up "Starting Directory Name".
Best,
Chao-Gan
回复: [RFMRI] Data Processing
已经解决了,谢谢您,以后我会仔细看视频的。
已经解决了,谢谢您,以后我会仔细看视频的。
DPARSF报错
Re: [RFMRI] Data Processing
这样啊,我从预处理开始就开着电脑,然后就睡觉去了
这样啊,我从预处理开始就开着电脑,然后就睡觉去了,早上起来就发现这样了,不过我后来接着重新运算了ALFF,就没有什么问题了,谢谢超赣兄!
ROI_OrderKey bugfix
Dear Chao-Gan,
I noticed a problem that the lastly defined ROI and it's ROI indices get lost when writing the ROI_OrderKey_*.csv files for the 'extract ROI signal' step. Could you please remove the following line at the bottom part of y_ExtractROISignal.m ?
MaskROILabel{1,length(ROIDef)} = []; % Force the undefined cells to emptyRegards,
Martin
Hi Martin,
Hi Martin,
Thanks a lot for your report!
I fixed it by changing into:
if size(MaskROILabel,2) < length(ROIDef) %YAN Chao-Gan, 131124. To avoid if the labels of the last ROI has been defined.
MaskROILabel{1,length(ROIDef)} = []; % Force the undefined cells to empty
end
This will be updated in the next version.
Thanks,
Chao-Gan
How read the rest fmri data of ADHD-200 with nii suffix in DPARS
Dear Mrs/Mr
Please, if possible, guide me how I can read the rest fmri data of ADHD-200 with nii suffix in DPARSF?
In your matlab based DPARSF, I can't load the my NIFTI file. Do I get the processed ADHD-200 competition data? I need the time course or BOLD signal for my research.
I will be grateful if you could help me.
Best regards,
Dear Hussain Montazery Kordy,
Dear Hussain Montazery Kordy,
I've met this problem recently and solved already. I post the solution here, hope to help you.
ADHD-200 data are 4-D, so you have to covert it to 3-D first.
funcdir='/public/home/FunImg';
cd ([funcdir '/']);
dirlist=dir(cd);
for j=3:length(dirlist)
restdir=[funcdir,filesep,dirlist(j).name,filesep];
cd (restdir);
PI='rest.nii';
PI=PI(1:end-3);
PO='.nii';
[Data, vox,Head] = rest_readfile(PI,'all');
if size(Data, 4)>1
for i=1:size(Data,4)
Index=['000',num2str(i)];
Index=Index(end-3:end);
[Path, fileN, extn] = fileparts(PO);
POout=[Path,filesep,fileN,'_',Index,extn];
rest_WriteNiftiImage(Data(:,:,:,i),Head,POout(end-7:end));
end
end
delete('rest.nii');
end
Best,
Jue WANG
dparsf
严老师,您好
我在使用dparsf处理数据,我先处理了一个人的数据是可以的,但是批处理时在normalize后老是报index ceceeds matrix dimensions的错误,请指教!
谢谢!
Hi, What's the parameter
Hi, What's the parameter setting and error message?
具体的参数设置和报错信息是什么呢?
超赣
Re: [RFMRI] Data Processing
All India Institute of Medical Sciences,
Ansari Nagar, New Delhi-110029
+919968569874
Hi Ansari,
Hi Ansari,
You can first select the template parameter of "VBM".
Then if you started with NIfTI images instead of DICOM images, change the starting directory name to the folder of "T1Img".
Best,
Chao-Gan
回复: [RFMRI] Data Processing
DPARSFA 2.3: Several fMRI sessions and one T1
comment deleted
??? Error using ==> y_ExtractROISignal at 188
FW: [RFMRI] Data Processing
Sent: Monday, December 07, 2015 7:41 PM
To: rfmri.org@rnet.co
Subject: Re: [RFMRI] Data Processing Assistant for Resting-State fMRI (DPARSF) V2.3
Correlation Analysis with text variable
Hi,
I want to do a correlation analysis between FC-Maps of a ROI and a text variable (duration of illness). What I did so far:
I use DPARSF --> statistical analysis --> Correlation analysis: Add FC-images of the patient-group (which are saved in the folder "FC-FunImgRWSDF) and add a text file with the respective duration of illness (in one column; I have 61 subjects, so 61 lines). The calculation works and I can open the result-file.
The first thing I am wondering about is that when I add the group images (so the folder "FC-FunImgRWSDF") the program immediatly reads in the standardized zFCmaps as well as the FCmaps. However I guess that I only want the zFCmaps to be included in the calculation, right? Do I just have to put the zFCmaps in a seperate folder and then add this folder as the image-folder or is there another way (like in one-sample-t-test, where I just type in a 0 for standardized and a 1 for unstandardized data).
Second, when I look at the result-image with a threshold of p=0.05 I see all significant (but uncorrected) correlations of the FCmap and the text-variable, am I right? If i now want to find out about the exact correlation of a respective cluster in this image, I can use the cluster-report. However this will only show me the correlation of the peak-voxel. Is there a way that the programm can tell me the overall correlation for the respective significant cluster?
Thansk in advance for the help :-)
Hi,
Hi,
1. "put the zFCmaps in a seperate folder and then add this folder as the image-folder"
2. If you want the mean correlation of the cluster. You can DPABI->Cluster->Save a single cluster to save an ROI file, and then use DPABI->Utilities->ROI Signals Extractor to extract it.
Best,
Chao-Gan
FC and zFCmaps
Thank you very much for the response. Another question: When I do a one-sample or two-sample t-test, do I always have to put the zFC maps in an extra folder? I thought that in a one-sample-t-test by setting the base to 0 (instead of 1) the program would automatically use just the zFC-maps.
Re: [RFMRI] Data Processing
individual zFCmaps
Thanks again :-)
I have another problem: I want to take a look at the individual zFCmaps. It works for almost every subject. However for 5 subjects I get the following error message once I try to open the zFCmaps in DPABI-Viewer:
Can you help me with this?
Best
Markus Loose
This is pretty weird. You can
This is pretty weird. You can upload the images to have a look.
FDR-Correction
Ok, the problem with viewing the single subject zFCmaps didn't occur again :-)
However I have another problem. I did a two-sample-t-Test of zFCmaps and now want to do an FDR-correction. I first opened DPABI-viewer, loaded up the T2-image and then just click in "FDR" and set the q-value to 0.05. Am I right?
I did that and unfortunately no differences were found. However a colleaque of mine is using the same data but a different programm and somehow he finds a group difference after FDR-correction. So I was wondering whether I might do it wrong in DPARSF. Also, the other programm filters the z-FCmaps on a threshold of 0.4 before doing any calculations such as t-tests. Only those voxels which are higher than the threshold of 0.4 will be included in the t-test, is that possible in DPARSF as well?
Greetings
Markus
Additional question
Another question: What threshold for the T-Value would you recommend when looking at the seed-network of one group (using one-sample-test of zFCmaps). For the Default-Mode-Netork for instance and a group-size of N=60 I use a T-value of 8 and the network looks quite good, however I am wondering whether this is the proper threshold for checking network quality.
thanks in advance :-)
Hi Markus,
Hi Markus,
You can perform one sample t-tests for each group, then combine the significant voxels across the two groups as a mask. This is achievable in DPABI.
For threshold in one sample t-test, you can perform GRF correction. Or some people set at p<1E-5 to get better visualization.
Best,
Chao-Gan
Hi Chao-Gan,
Hi Chao-Gan,
thanks for the reponse :-)
It sounds very interesting, however, I am not sure whether I am doing it right. How do I create a mask upon the significant voxels of both one-sample-t-tests? I tried uploaing both t-test images (of patients and controls) in the DPABI-Viewer, then I set the threshold to 0.000001 and then clicked on save all clusters. This will produce a mask which I then uploaded as a mask when doing the FDR-Correction. Is that the procedure you were talking about or should I do it differently?
Also, which parameters should I use in a GRF-correction. Would you recommend both p=0.05 in Voxel and cluster p-value?
Greetings from Germany
Markus
Re: [RFMRI] Data Processing
Deputy Director, MRI Research Center
Institute of Psychology, Chinese Academy of Sciences
16 Lincui Road, Chaoyang District, Beijing 100101, China
Allright, thanks :-)
Allright, thanks :-)
When I load up the images in the image calculator, do I have to add anything under "Group images"?
I then do FDR correction with the two-sample-t-test and load up the mask in the FDR-correction-window, is that right?
Also, when doing GRF correction, what p-value for cluster respectively voxel would you recommend when using it in a one-sample-t-test to check for network quality. Do I have to load up a mask there as well?
Best,
Markus
Re: [RFMRI] Data Processing
Deputy Director, MRI Research Center
Institute of Psychology, Chinese Academy of Sciences
16 Lincui Road, Chaoyang District, Beijing 100101, China
Additional FDR?
Ok, thanks that was helpful :-)
When I view the one-sample-t-test for a network of one group and set the threshold to p<0.000001 should I do an additional FDR correction or is that unnecessary since the p-value is that small?
Also, when I do an GRF-correction on a one-sample-t-test to check network quality, do I have to upload a mask as well?
1. I think that's OK without
1. I think that's OK without further FDR correction.
2. Yes, you need to apply a mask.
Best,
Chao-Gan
老師
老師
我使用matlab 2016, spm8, DPARSF_V2.3_130615 run前置分析
但是一直都在FunImgAR此步驟出錯,出錯畫面如下
Please try the newest version
Please try the newest version of DPABI/DPARSF.
预处理顺序